UGene (BioInformatics)
Overview and Setup
UGENE is a bioinformatics software toolkit that provides a user-friendly environment for managing, analyzing, and visualizing biological data. It integrates a wide range of established bioinformatics tools for tasks like genome sequence analysis, Next Generation Sequencing (NGS) data processing, multiple sequence alignment, and phylogenetic tree construction.
To access the OOD web portal, please refer to our Getting Started with Open OnDemand documentation page. Once logged into the portal, you can launch UGENE from under the Bioinformatics header.
Configuring UGENE
To get started, connect to OOD and choose UGENE from the Bioinformatics header.
In the submission form, you can choose the account, partition, the length of your job, the number of cores you need, and the node type.
Note
- For account, put whatever group account you’re associated with. If you do not have a group account, use free.
- The partition is always ood
- Node type is any
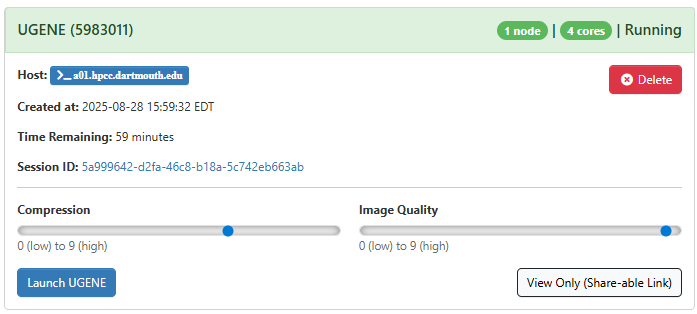
After you click “Launch”, a new session will be queued. Once the session is active your page will appear as follows:
When launching UGENE, you’ll notice two key sliders: compression and image quality. These settings can greatly affect your overall experience. To optimize performance, we suggest cranking up the image quality slider to the highest setting (9). If you’re not satisfied with the result, it’s easy to close the tab, adjust the sliders, and start again. Experimentation is key to finding the perfect balance between performance and visual quality for your environment.
Using UGENE
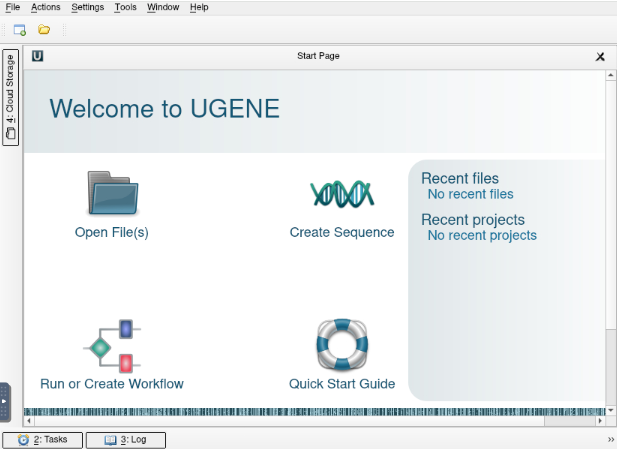
Click on “Launch UGENE” to begin using UGENE. Once you have entered your session, you should see the UGENE page below:
This is a typical UGENE interface that should be familiar to frequent users.